SEEDs enable enrichment of cells with transgene integrations
To develop a method for cells with transgene integrations to be enriched through negative selection, we designed two reagents: (1) a guide RNA (gRNA) targeting an intron of a surface-expressed protein that generates a DSB that minimally impacts expression and (2) a SEED HDR template (HDRT) that uses synthetic splice acceptor (SA) and splice donor (SD) sequences to introduce an in-frame transgene payload preceded by a P2A sequence at a position that disrupts the target protein (Fig. 1a).
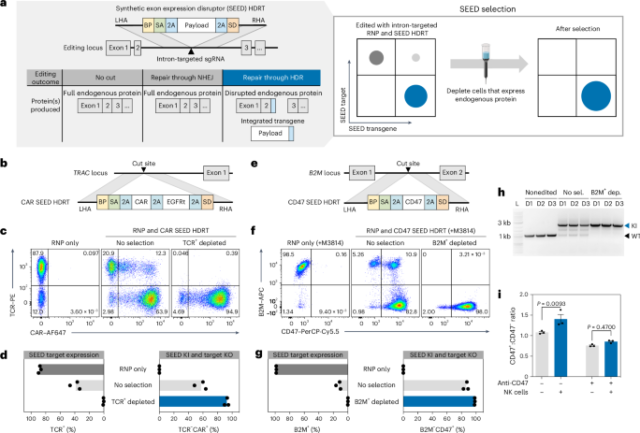
a, Overview of editing outcomes generated with an intron-targeted single gRNA (sgRNA) and a SEED HDRT. Immunomagnetic reagents are used to deplete cells that retain expression of the surface protein targeted by a SEED, thereby enriching for cells with transgene integration. b, Diagram of a TRAC intron-targeted SEED HDRT encoding a CAR and EGFRt. c,d, T cells were edited with TRAC intron-targeted RNP and HDRT (b) and then immunomagnetically purified with anti-TCR (n = 3 donors). c, Flow cytometry plots of TCR and CAR expression (anti-mouse F(ab′)2). d, Percentage of TCR+ and TCR− CAR+ cells. KI, knock-in; KO, knockout. e, Diagram of a B2M intron-targeted SEED HDRT encoding CD47. f–h T cells were edited with B2M intron-targeted RNP and HDRT (e) in the presence of M3814 then immunomagnetically purified with anti-B2M (n = 3 donors). f, Flow cytometry plots of B2M and CD47 expression. g, Percentage of B2M+ and B2M– CD47+ cells. h, Genomic DNA PCR targeting the SEED integration site at B2M. Amplicon for nonedited alleles, black triangle; amplicon for HDRT integration, blue triangle (n = 3 donors). No sel.; no selection; B2M+ dep.; B2M+ depleted. i, Fold enrichment of CD47+ SEED-edited cells relative to CD47− cells after coculture with NK cells (n = 3 T cell donors). Data are displayed as the mean ± s.e.m. Significance was assessed with a two-way analysis of variance (ANOVA) and Šidák’s multiple-comparisons test. LHA, left homology arm; RHA, right homology arms. APC, allophycocyanin; PE, phycoerythrin.
We initially targeted the two most common loci for therapeutic T cell engineering, TRAC and B2M (β2 microglobulin). Insertion of a chimeric antigen receptor (CAR) or transgenic TCR at the TRAC locus can redirect T cell specificity and benefits from endogenous regulatory elements to produce potent and durable T cell therapies23,24. B2M is required for major histocompatibility complex class I (MHC-I) expression and can be disrupted to help evade host T cell recognition in allogeneic settings25. To identify optimal SEED integration sites for TRAC and B2M, we designed two panels of intron-targeted gRNAs and screened for guides that generate indels in primary human T cells without disrupting TCR or B2M surface expression, respectively (Extended Data Fig. 2a–d). Candidate gRNAs exhibited high indel generation efficiencies (up to 97%) with minimal disruption of TCR (9–16%) or B2M (1–2%) surface expression.
To confirm that SEED integration could disrupt TCR surface expression, we edited T cells with a TRAC intron-targeted ribonucleoprotein (RNP) and an adeno-associated viral vector (AAV6) encoding a SEED HDRT for a CD19-specific CAR and a truncated epidermal growth factor receptor (EGFRt) tag (Fig. 1b,c)18. Integration-mediated TCR disruption (TCR−CAR+) occurred in over half (57.4%) of SEED-transduced cells. SEED-transduced cells also exhibited less integration-independent TCR disruption (TCR−CAR−) than cells edited with the RNP alone, in agreement with reports that template-mediated HDR can outcompete pathways that create large deletions (Fig. 1c and Extended Data Fig. 3a)26. Although most T cells express one TCRα chain, a subset of TCR+CAR+ cells was also observed (Fig. 1c), which could result from monoallelic HDRT integration in the ~30% of T cells that exhibit transcription at both TCRα alleles27. Immunomagnetic TCR depletion removed nearly all TCR+ cells and recovered a highly pure (92.6%) TCR−CAR+ population with yields of up to 76% (Fig. 1c,d and Extended Data Fig. 4).
To compare editing outcomes between SEED and exon-targeting strategies, we designed and tested an analogous CAR HDRT with an integration site in TRAC exon 1 (Extended Data Fig. 3b,c)18. CAR expression and homogeneity were similar in T cells edited with both constructs (Extended Data Fig. 3d,e). However, immunomagnetic TCR depletion of cells edited with an exon-targeted HDRT minimally enriched for TCR−CAR+ cells, as the majority of CAR− cells were also TCR− (Extended Data Fig. 3f,g). These data demonstrate that SEEDs enable edited cells to be enriched by coupling transgene integration to target protein disruption.
NHEJ inhibitors enable efficient biallelic SEED integration
The TCR is subject to multiple pretranslational and posttranslational processes that result in functional allelic exclusion and editing at the single active TCRα allele is often sufficient to ablate TCR expression23,27. In contrast, editing at both alleles is necessary to fully disrupt B2M and most other loci. To identify the editing conditions necessary for biallelic SEED integration, we tested an AAV6 SEED HDRT designed to disrupt B2M expression and deliver CD47, an immune checkpoint molecule that has been shown to inhibit natural killer (NK) and macrophage activity against cells lacking MHC-I expression (Fig. 1e)28,29,30.
We characterized editing outcomes in T cells transduced at multiplicities of infection (MOIs) ranging from 1 × 103 to 3 × 105 (Extended Data Fig. 5a–e). In a subset of conditions, editing was performed in the presence of M3814, an NHEJ inhibitor that promotes biallelic transgene integration31. After editing, distinct B2M+CD47+ and B2M−CD47+ populations were observed (Extended Data Fig. 5a). B2M+CD47+ cells expressed intermediate levels of B2M and CD47 (corresponding to monoallelic SEED integration), while B2M−CD47+ cells exhibited loss of B2M expression and increased CD47 expression (corresponding to biallelic SEED integration) (Extended Data Fig. 5a,b).
Biallelic SEED integration was achieved in 83% of cells following transduction at the highest MOI and with M3814 (Extended Data Fig. 5d). As expected, biallelic SEED integration was reduced when lower MOIs were used or when editing was performed without M3814 (Extended Data Fig. 5d,e)31. However, biallelic integration was still detectable in the least efficient editing conditions (lowest MOI), where the total fraction of CD47+ cells was <10% (Fig. 5c–e). These data suggest that optimized editing conditions enable biallelic integration to become the primary editing outcome but are not required for biallelic integration to occur.
SEED-Selection isolates cells with biallelic integrations
To test whether SEED-Selection could enable the isolation of cells with biallelic transgene integrations, we performed immunomagnetic B2M depletion after editing cells with a B2M-targeted SEED HDRT (Fig. 1f,g). Immunomagnetic selection efficiently depleted B2M+CD47− and B2M+CD47+ cells and enabled the recovery of a highly pure (>98%) population of cells with biallelic SEED integration (B2M−CD47+), with yields of up to 56% (Fig. 1f,g and Extended Data Fig. 4). Product purity was further evaluated through genomic DNA PCR, which confirmed depletion of nonedited B2M alleles during selection (Fig. 1h).
To assess the performance of SEED-Selection in samples with different editing distributions, we immunomagnetically purified samples transduced with high (3 × 105) or low (1 × 104) MOIs (Extended Data Fig. 5f,g). While the final purity of B2M−CD47+ cells was higher in samples transduced with a high MOI, SEED-Selection resulted in a larger improvement in B2M−CD47+ purity in samples transduced with a low MOI (Extended Data Fig. 5f,g). These data demonstrate the versatility of SEED-Selection across different editing scenarios.
To test whether CD47 expression was sufficient to reduce NK cell cytotoxicity, we generated a mixture of T cells with two subsets (B2M−CD47+ and B2M−CD47−) and performed a coculture with activated primary human NK cells. The composition of the coculture was then quantified by flow cytometry and compared to that of T cells alone. As intended, B2M−CD47+ cells were enriched relative to B2M−CD47− cells after coculture (Fig. 1i and Extended Data Fig. 5h). B2M−CD47+ enrichment was not observed when T cells were pretreated with a CD47-blocking antibody, confirming that SEED-mediated CD47 activity facilitated NK cell evasion (Fig. 1i and Extended Data Fig. 5h).
SEED-Selection simultaneously enriches for multilocus edits
To test whether SEED integration could be achieved at multiple loci simultaneously, we edited T cells with both TRAC-CAR and B2M-CD47 SEEDs (Fig. 2a–c). Cells were transduced at an MOI of 2 × 105, an AAV dose chosen to mimic preclinical studies for CaMMouflage (NCT05722418), an ongoing clinical trial evaluating CRISPR-edited T cells with integrations at TRAC and B2M (ref. 32). High integration efficiencies were observed at both target loci with up to 55% of cells undergoing full editing (TCR−CAR+B2M−CD47+), confirming that multilocus integration could be attained with clinically relevant doses of AAV (Fig. 2b,c). Simultaneous immunomagnetic depletion with anti-TCR and anti-B2M antibodies further enriched for cells with transgene integrations, allowing for the isolation of a highly pure (89.4%) population of fully edited cells (Fig. 2b,c).
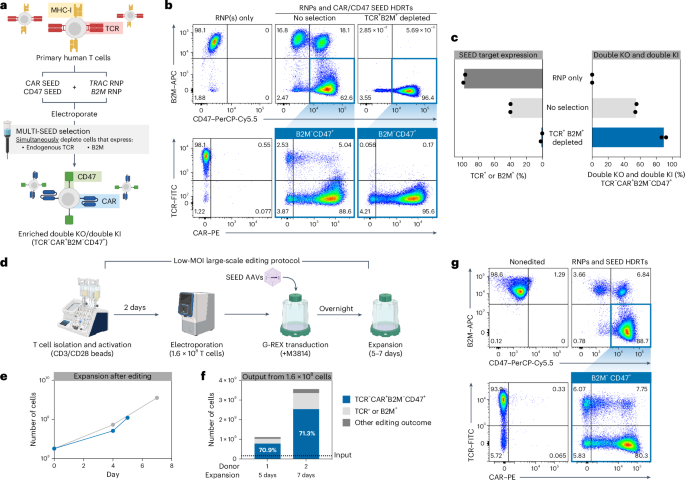
a–c, T cells were edited with TRAC and B2M RNPs (+M3814) and transduced with TRAC-CAR SEED and B2M-CD47 SEED HDRTs. Edited cells were then immunomagnetically purified with anti-B2M and anti-TCR (n = 2 donors). a, Multiplexed editing and enrichment strategy. b, Representative flow cytometry plots of TCR, CAR, B2M and CD47 expression. c, Percentage of B2M+ or TCR+ cells and fully edited cells (TCR−CAR+B2M−CD47+). –d–g 1.6 × 108 T cells were edited with TRAC and B2M RNPs and transduced with TRAC-CAR SEED and B2M-CD47 SEED HDRTs using a GMP-compatible workflow (n = 2 donors). d, Overview of large-scale editing workflow. e, Expansion of cells after multiplexed editing. f, Number and composition of edited cells after expansion. g, Representative flow cytometry plots of TCR, CAR, B2M and CD47 expression. FITC, fluorescein isothiocyanate.
To improve our ability to produce complex cell therapies at scale, we designed an optimized protocol using G-REX flasks on the basis of reports showing that AAV transduction is influenced by culture density (Extended Data Fig. 6a)33. Using this improved workflow, we were able to achieve complete editing at TRAC and B2M in ~50% of cells using 85% less virus (MOI of 3 × 104) than our original protocol (Extended Data Fig. 6b,c). Robust expansion was observed following editing, enabling up to 1.6 × 107 fully edited T cells to be generated from an initial input of 4 × 106 cells after a 7-day expansion (Extended Data Fig. 6d). Multiplexed immunomagnetic depletion after expansion increased the fraction of fully edited cells from 50% to 81% with yields of up to 77% (Extended Data Fig. 4 and Extended Data Fig. 6b,c). These data demonstrate that SEED-Selection can be multiplexed to isolate homogenous cell products that have multiple desirable integrations.
Multilocus low-MOI editing is efficient at clinical scale
To validate the scalability of our optimized editing protocol, we performed two clinical-scale manufacturing runs editing TRAC and B2M in 1.6 × 108 primary human T cells using good manufacturing practice (GMP)-compatible reagents, equipment and processes (Fig. 2d). Complete editing was achieved in 71% of cells, with outputs of up to 2.5 × 109 fully edited cells after only 7 days of expansion, indicating that multilocus integration strategies can be efficiently implemented in clinical settings (Fig. 2d–g).
Epitope-edited human leukocyte antigen-independent TCRs are compatible with SEED-Selection
Synthetic immune receptors containing TRAC and TRBC domains, such as transgenic α/β TCRs34 or human leukocyte antigen (HLA)-independent TCRs (HITs)35, are highly sensitive to low antigen densities. However, these receptors are incompatible with TCR SEED-Selection as they are also bound by the anti-TCRα/β antibodies used to deplete nonmodified and partially edited cells (Fig. 3a–c). Structural analysis and high-throughput screening techniques have been used to design epitope-edited receptors that evade a specific antibody and remain functional36,37,38. We hypothesized that epitope editing of SEED payloads would allow for the enrichment of cells with a transgene that would otherwise be depleted in SEED-Selection.
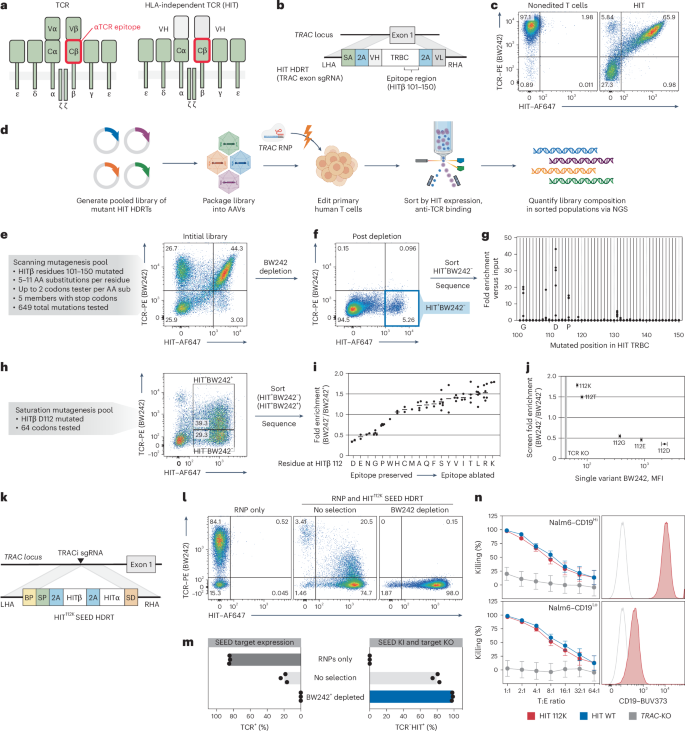
a, Diagram of a TCR and a HIT. b, Diagram of a TRAC exon-targeted HDRT encoding a HIT. c, Flow cytometry of anti-TCR (BW242) binding and HIT expression (anti-mouse F(ab′)2) in T cells with a HIT at TRAC (n = 1 donor). d, Screening workflow for epitope mapping. e–g T cells were edited to express a library of mutated HITs then purified with BW242. BW242 binding and HIT expression were quantified with flow cytometry before (e) and after (f) purification. HIT+ BW242– cells were sorted (blue box) for sequencing (n = 1 donor). g, Enrichment of mutations in HIT+BW242− cells (blue box in f) compared to the original library (e). Dots represent the average of tested codons for an amino acid. h, Flow cytometry of BW242 binding and HIT expression in T cells edited with a HIT β112 saturation mutagenesis pool. Boxes indicate sorted populations. i, Enrichment of mutations in HIT+BW242− cells versus HIT+BW242+ cells. Dots represents enrichment for a single codon (n = 1 donor). Lines display the average enrichment for an amino acid. j, BW242 binding for HIT+ T cells edited with HIT variants (n = 3 donors) compared with average enrichment scores (from i). The black line indicates the BW242 median fluorescence intensity (MFI) for TCR– cells. The MFI s.e.m. is represented by bars (HIT mutants) or the shaded gray area (TCR–). k, Diagram of a TRAC SEED HDRT encoding an epitope-edited HIT. l,m, T cells were edited with a HIT SEED (+M3814) and then purified with BW242 (n = 3 donors) l, Flow cytometry of BW242 binding and HIT expression m, Percentage of BW242+ and BW242−HIT+ cells. n, Cytotoxic activity of T cells with a nonmodified HIT (blue), epitope-edited HIT (red) or TCR knockout (gray) against Nalm6 lines (technical triplicate; data shown from one of four donors). Histograms show Nalm6 CD19 expression (flow cytometry). Unstained cells are shown in gray. Data are displayed as the mean ± s.e.m. T, target; E, effector.
Previous studies established that certain anti-TCRα/β antibody epitopes can be disrupted by murinizing a portion of the β constant domain (Cβ) of a TCR24,39. To identify single-amino-acid substitutions capable of disrupting antibody binding, we systematically substituted individual residues across a 50-aa span of the HIT Cβ in the form of a 649-member pooled knock-in library (Fig. 3d,e). After introducing the receptor pool into the TRAC locus of T cells, we immunomagnetically depleted cells that were bound by a GMP-grade anti-TCRα/β antibody (clone BW242/412; hereafter referred to as BW242) and flow-sorted for cells that retained high HIT expression (Fig. 3f). Library member abundance was then quantified by RNA-based next-generation sequencing (NGS) (Fig. 3g)8. While most library members were depleted, a subset of substitutions at Cβ residues G102, D112 or P116 were enriched, suggesting that these positions interact with BW242 and can be substituted to prevent HIT receptor depletion without compromising surface expression.
To catalog conservative and nonconservative substitutions at these key positions, we generated three pooled libraries of HIT receptors with saturation mutagenesis performed at Cβ residues 102, 112, or 116. Each library was individually introduced into T cells and HIT+ cells were flow-sorted on the basis of BW242 binding (Fig. 3h and Extended Data Fig. 7a). RNA-based NGS was then used to assess the relative enrichment of mutants in each bin as compared to the original library. Residues at each position were ranked from least to most conservative according to the ratio of enrichment between bins (Fig. 3i and Extended Data Fig. 7b). As expected, the native Cβ residue (G102, D112 and P116) at each position was the most conservative. Almost all substitutions at 102 and 116 were preferentially enriched in the BW242− bin, suggesting that the native residues at these positions are required for optimal BW242 binding (Extended Data Fig. 7b). In total, 14 of 19 substitutions at Cβ residue 112 were also enriched in the BW242– bin (Fig. 3i). However, substitutions to E, N, G, P and W were preferentially enriched in the BW242+ bin. The most conservative substitution at Cβ residue 112 preserved the charge (D to E) whereas the least conservative ones switched the charge (D to K or D to R), suggesting that electrostatic interactions with D112 contribute to BW242 binding.
To confirm that this sequencing-based approach for epitope mapping predicts changes in antibody binding, we generated five HIT receptors with different amino acids at Cβ residue 112 and individually introduced them into T cells. BW242 binding was assessed by flow cytometry and compared to the enrichment ratios obtained from the Cβ residue 112 saturation mutagenesis screen (Fig. 3j and Extended Data Fig. 7c). Sequencing-based epitope mapping correctly ranked the panel of substitutions on the basis of epitope conservation and accurately detected variation in BW242 binding across the dynamic range. HIT and CD3 surface expression was maintained across receptor designs, confirming that variations in BW242 binding were not because of alterations in HIT expression or assembly with CD3 chains (Extended Data Fig. 7d). Of the five tested mutants, HIT K112 had the least BW242 binding and was selected for functional characterization (Fig. 3j).
We introduced a CD19-specific HITK112 into a TRAC-targeted SEED and immunomagnetically purified edited T cells. Depletion with BW242 resulted in complete removal of cells with endogenous TCRs and allowed for the recovery of a pure (>98%) population of HIT+ cells with yields of up to 68% (Fig. 3k–m and Extended Data Fig. 4). As the HIT receptor was developed to have the capacity to target tumor cells expressing low antigen densities35, we assessed receptor function through cytotoxicity assays with target cell lines expressing high or low levels of CD19. HITK112 performed similarly to nonmodified HIT (HITWT) against lines with high and low antigen density, confirming that epitope editing allows HIT+ cells to be enriched through SEED-Selection without compromising receptor function (Fig. 3n and Extended Data Fig. 7e).
Epitope-edited TCRs enable minimization of mispairing
T cells can be engineered to target a specified peptide-MHC through the expression of a transgenic TCR34. However, mispairing between endogenous and transgenic TCR chains can occur when a transgenic TCR is expressed in an otherwise nonedited T cell24,40. Because all TCR chains compete for the same pool of CD3 molecules during assembly, mispairing can reduce the surface expression of the correctly paired transgenic TCR24,40. Mispairing-associated autoreactivity has also been reported in mouse models but the clinical importance of this phenomenon has not yet been established40.
Mispairing is a nonrandom process influenced by properties of the transgenic TCR and each T cell’s endogenous TCR and some combinations of TCR chains pair inefficiently41. In TRAC-edited cells engineered to express a transgenic TCR, mispairing occurs between the transgenic TCRα and endogenous TCRβ (ref. 24). Therefore, we hypothesized that editing the BW242 epitope on a transgenic TCRβ chain could allow cells that express a correctly paired transgenic TCR (which should evade BW242 binding) to be distinguished from cells that express a mispaired TCR (which should retain BW242 binding) (Fig. 4a).
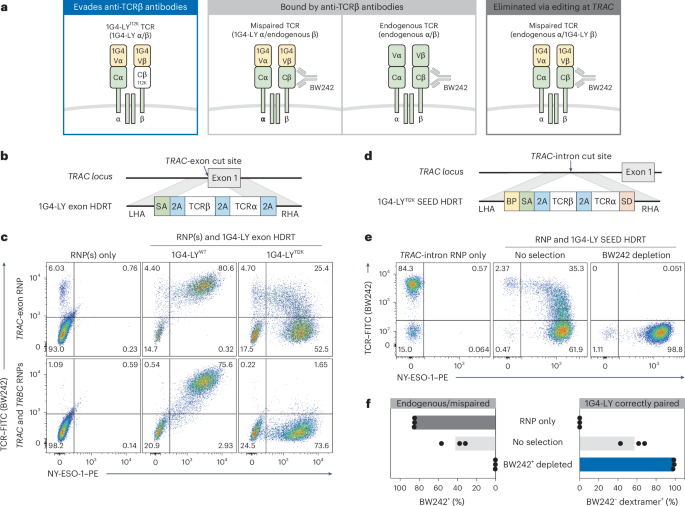
a, Schematic of possible TCR pairs in nonedited and TRAC-edited T cells engineered to express the 1G4-LYK112 TCR (specific for NY-ESO-1). b, Diagram of a TRAC exon-targeted HDRT encoding 1G4-LY. c, Flow cytometry plots of BW242 binding and NY-ESO-1 dextramer binding in CD8+ cells. Editing was performed with RNPs targeting TRAC or TRAC/TRBC and HDRTs (a) encoding nonmodified (1G4-LYWT) or epitope-edited (1G4-LYK112) versions of 1G4-LY (n = 2 donors). d, Diagram of a TRAC intron-targeted SEED HDRT encoding 1G4-LY. e,f, T cells were edited with a 1G4-LY SEED (+M3814) (d) and immunomagnetically purified with BW242 (n = 3 donors). e, Flow cytometry plots of BW242 binding and NY-ESO-1 dextramer binding for cells gated on CD8+. f, Percentage of cells with an endogenous or mispaired TCRs (BW242+) or correctly paired 1G4-LY (dextramer+BW242–).
To test this hypothesis, we generated an epitope-edited variant of 1G4-LY: a clinically validated, affinity-matured, transgenic TCR that targets NY-ESO-1 (SLLMWITQC) on HLA-A*02 (ref. 42). We designed TRAC exon-targeted HDRTs to simultaneously disrupt the endogenous TCRα and facilitate expression of a nonmodified 1G4-LY (1G4-LYWT) or an epitope-edited variant (1G4-LYK112) (Fig. 4b). Each HDRT was introduced into CD8+ T cells edited with an RNP targeting TRAC. Multiplexed editing of TRAC and TRBC (TCRβ constant) was also performed to produce T cells where 1G4-LY was expressed and both endogenous TCR chains were disrupted, ensuring exclusive expression of correctly paired 1G4-LY.
NY-ESO-1 dextramer binding correlated with BW242 binding in cells expressing 1G4-LYWT, as expected (Fig. 4c). In contrast, TRAC-edited cells engineered to express 1G4-LYK112 exhibited a distinct population with correct TCR pairing (dextramer+BW242−) and a separate population with apparent TCR mispairing (dextramer+BW242+). In agreement, this population was eliminated by concurrent knockout of the endogenous TCRβ chain (Fig. 4c). Dextramer+ cells in all conditions expressed similar levels of CD3, confirming that variations in BW242 binding were not caused by differences in TCR assembly (Extended Data Fig. 8a). We also used epitope editing to identify and deplete mispaired TRAC-edited cells engineered to express a transgenic TCR (DMF5) targeting MART1 (melanoma antigen recognized by T cells 1), supporting the generalizability of this approach (Extended Data Fig. 8b,c)43.
To test whether an epitope-edited TCR could be enriched through SEED-Selection, we introduced 1G4-LYK112 into a TRAC-targeted SEED and edited CD3+ T cells (Fig. 4d). Depletion with BW242 resulted in efficient removal of cells with endogenous or mispaired TCRs and the recovery of a highly pure (>98%) population of correctly paired 1G4-LY+ cells with yields of up to 60% (Fig. 4e,f and Extended Data Fig. 4). This result suggests that SEED-Selection could be used to deplete T cells with undesired specificities without the need to perform simultaneous editing at TRAC and TRBC.
SEED enables simultaneous coreceptor and TCR swapping
Recent studies have emphasized the role of CD4+ T cells as contributors to long-lasting immune responses to tumors44. However, the MHC-I-restricted TCRs commonly isolated for transgenic TCR therapies perform suboptimally in the absence of the CD8α/β coreceptor45,46,47. Although 1G4-LY has been demonstrated to undergo coreceptor-independent activation42, we observed that dextramer binding was markedly reduced in edited CD4+ T cells as compared to edited CD8+ T cells (Extended Data Fig. 9a). Therefore, we sought to develop a strategy for isolating CD4+ T cells edited to express both CD8α/β and 1G4-LY.
As CD4 should be superfluous in cells engineered to express an MHC-I-restricted TCR, we screened for nondisruptive gRNAs targeting the CD4 locus and designed a SEED HDRT to co-deliver CD8α and CD8β (Fig. 5a and Extended Data Fig. 9b). Integration-mediated CD4 disruption (CD4−CD8+) was achieved in >75% of HDRT-transduced CD4+ cells (Fig. 5b,c). Full disruption of CD4 requires both alleles to be nonfunctional. Correspondingly, a population that coexpressed intermediate levels of CD4 and CD8 was observed after editing, consistent with monoallelic SEED integration (Extended Data Fig. 9c). Immunomagnetic CD4 depletion allowed for the recovery of a pure (>98%) population of cells with biallelic SEED integration (CD4−CD8+) with yields of up to 79% (Fig. 5b,c and Extended Data Fig. 4).
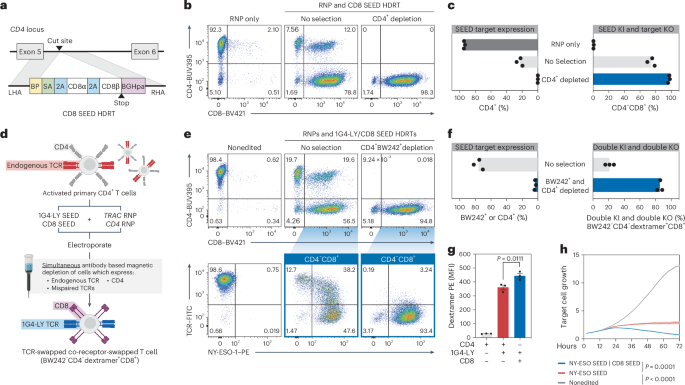
a, Diagram of a CD4 intron-targeted SEED HDRT encoding CD8α/β. b,c, CD4+ T cells were edited with a CD8 SEED (a) (+M3814) and immunomagnetically purified with anti-CD4 (n = 3 donors). b, Flow cytometry plots of CD4 and CD8 expression. c, Percentage of CD4+ and CD4−CD8+ cells. d–h CD4+ T cells were edited with 1G4-LY and CD8 SEEDs (+M3814) and immunomagnetically purified with BW242 and anti-CD4 (n = 3 donors). d, Overview of editing and selection strategy. e, Flow cytometry plots of CD4 and CD8 expression and BW242 and NY-ESO-1 dextramer binding. f, Percentage of cells that express CD4 or an endogenous or mispaired TCR (BW242+). Percentage of fully edited cells with minimal mispairing (BW242−CD4−dextramer+CD8+). g, Assessment of NY-ESO-1 dextramer binding by flow cytometry in CD4+CD8−BW242−1G4-LY+ cells (red), CD4−CD8+BW242−1G4-LY+ cells (blue) and nonedited CD4+ T cells (gray) (n = 3 donors). Data are displayed as the mean ± s.e.m. h, Assessment of cytotoxic activity of CD4+ T cells against NY-ESO-1+ A375 target cells (technical triplicate; data shown from one of four donors). Nonedited T cells, dark gray; 1G4-LY SEED-edited, red; 1G4-LY SEED-edited; CD8 SEED-edited, blue. Data are displayed as the mean (bold line) ± s.e.m. (shaded area). Significance was assessed at 72 h. The significance in g,h was assessed using a repeated-measures one-way ANOVA and Turkey’s multiple-comparisons test.
To assess whether the performance of 1G4-LY in CD4+ cells was improved through overexpression of CD8α/β, we edited T cells with a TRAC-targeted 1G4-LYK112 SEED alone or with a CD4-targeted CD8α/β SEED (Fig. 5d). Each cell population was immunomagnetically purified with anti-TCR (BW242) and anti-CD4 antibodies to enrich for transgene integration and to deplete cells with mispaired and endogenous TCRs (Fig. 5d–f and Extended Data Fig. 4). As expected, cells that expressed both 1G4-LYK112 and CD8α/β exhibited increased NY-ESO-1 dextramer binding in comparison to cells that expressed 1G4-LYK112 alone (Fig. 5g). Furthermore, in a longitudinal cytotoxicity assay, coexpression of CD8α/β improved the control of A375, a melanoma cell line that endogenously expresses NY-ESO-1 (Fig. 5h and Extended Data Fig. 9d).
To validate the ability of SEED-Selection to isolate fully edited cells after complex editing, we sought to simultaneously select for cells with clinically desirable transgenes integrated at three loci (Fig. 6a). CD4+ cells were edited with RNPs targeting TRAC, B2M and CD4 and transduced with SEEDs encoding 1G4-LYK112, CD47 and CD8. After editing, cells expressing any combination of endogenous TCR, mispaired TCR, B2M and CD4 were immunomagnetically removed in a single step (Fig. 6b,c). Depletion of all target markers was efficient, resulting in the recovery of highly pure (up to 90%) populations of fully edited BW242−B2M−CD4−1G4-LY+CD47+CD8+ cells with yields of up to 50% (Fig. 6b,c and Extended Data Fig. 4). Multiplexed SEED-Selection was also performed on cells transduced with our low-MOI protocol, where immunomagnetic selection increased the percentage of fully edited nonmispaired cells from 24% to 70% (Extended Data Fig. 4 and Extended Data Fig. 10a–c).
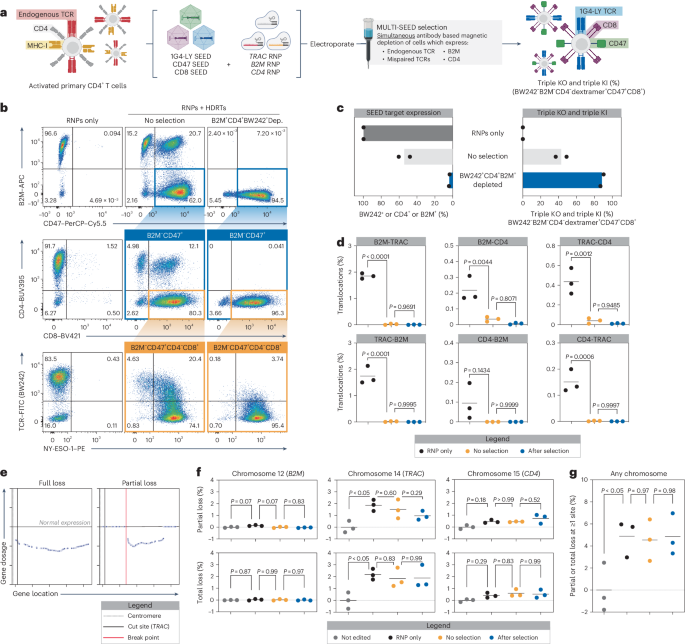
a–c, CD4+ T cells were edited with RNPs targeting TRAC, B2M and CD4 (+M3814) and transduced with SEEDs encoding 1G4-LY, CD8 and CD47, respectively. Edited cells were then immunomagnetically purified with BW242, anti-CD4 and anti-B2M (n = 2 donors). a, Diagram of the workflow for multiplexed editing and enrichment. b, Flow cytometry plots of SEED target expression (endogenous TCR, CD4 and B2M), SEED payload expression (CD8 and CD47) and NY-ESO-1 dextramer binding. c, Percentage of cells expressing any SEED target, a mispaired TCR (BW242+, CD4+ or B2M+) or triple-knockout or triple-knock-in cells with correct 1G4-LY pairing (BW242−B2M−CD4−dextramer+CD47+CD8+). d, Assessment of balanced translocations among TRAC, B2M and CD4 after multiplexed editing (+M3814) and SEED-Selection by ddPCR after a 9-day expansion (n = 3 donors). e–g, partial and total chromosome loss was assessed in after multiplexed editing and SEED-Selection by single-cell RNA sequencing (n = 3 donors). e, Representative gene expression profiles of cells with partial or complete loss of chromosome 14 after multiplexed editing. f, Normalized frequency of cells with partial and total chromosomal loss. g, Normalized frequency of cells with one or more chromosomal abnormalities. The significance in d,f,g was assessed using a repeated-measures one-way ANOVA and Turkey’s multiple-comparisons test.
Evaluation of genome stability after multilocus SEED editing
Rearrangements such as translocations and chromosome loss have been observed in gene-edited primary cells48,49,50. To understand how SEED-Selection influences translocation frequency, we designed an array of digital droplet PCR (ddPCR) assays and quantified balanced translocations 9 days after simultaneous editing at TRAC, B2M and CD4. When editing was performed in the absence of HDRTs, individual translocation frequencies ranged from 0.1% to 1.9% (Fig. 6d). As previously observed, translocation frequencies decreased significantly when editing was performed with HDRTs (Fig. 6d)51. Translocations could hypothetically result in integration-independent disruption of SEED target expression. However, further isolation of fully edited cells by SEED-Selection did not significantly alter translocation frequencies, suggesting that cells with translocations may often retain expression of at least one SEED target (Fig. 6d). These data build on previous findings that unintended editing outcomes can be prevented through the addition of an HDRT and validate that SEED-Selection does not enrich for balanced translocations.
To determine whether SEED-Selection enriches for genome rearrangements that result in integration-independent SEED target disruption through partial or complete chromosome loss, we used a previously described single-cell RNA sequencing-based workflow to evaluate the frequency of these events at chromosome 12 (B2M), chromosome 14 (TRAC) and chromosome 15 (CD4) in comparison to a nonedited control (Fig. 6e). Both partial and complete chromosome loss in HDRT edited cells was more frequent at chromosome 14 (1.5% partial and 1.8% total) versus chromosome 12 (<0.01% partial and 0.02% total) or chromosome 15 (0.5% partial and 0.6% total), which might be because of the proximity of the TRAC locus to the centromere, correlating with the increased frequency of integration-independent SEED target disruption observed with TRAC gRNAs (Fig. 6e,f). The occurrence of partial and complete loss of these chromosomes was not significantly affected by the presence of an HDRT during editing or by SEED-Selection (Fig. 6f). To globally evaluate changes in genome integrity, we also quantified the frequency of cells with at least one partial or total deletion involving any autosomal chromosome and found no significant differences associated with SEED-Selection (Fig. 6g). These results further validate that large-scale genomic rearrangements involving target loci are not enriched through SEED-Selection.